By combining multiple statistical techniques we created MLRP. In this post we will give a short overview of the technique by applying it to the iris data set (which can be found here), which is one of the best known data sets found in the classification and pattern recognition literature. The data consists of 150 observations of iris plants, which were measured in terms of sepal length (SL), sepal width (SW), petal length (PL), and petal width (PW).
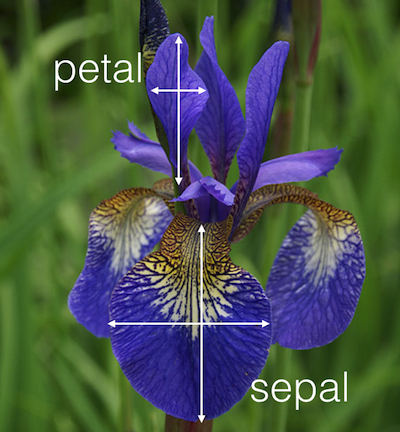

The outcome classes are species of iris plants: Setosa (sets), Versicolour (vrsc), and Virginica (vrgn).

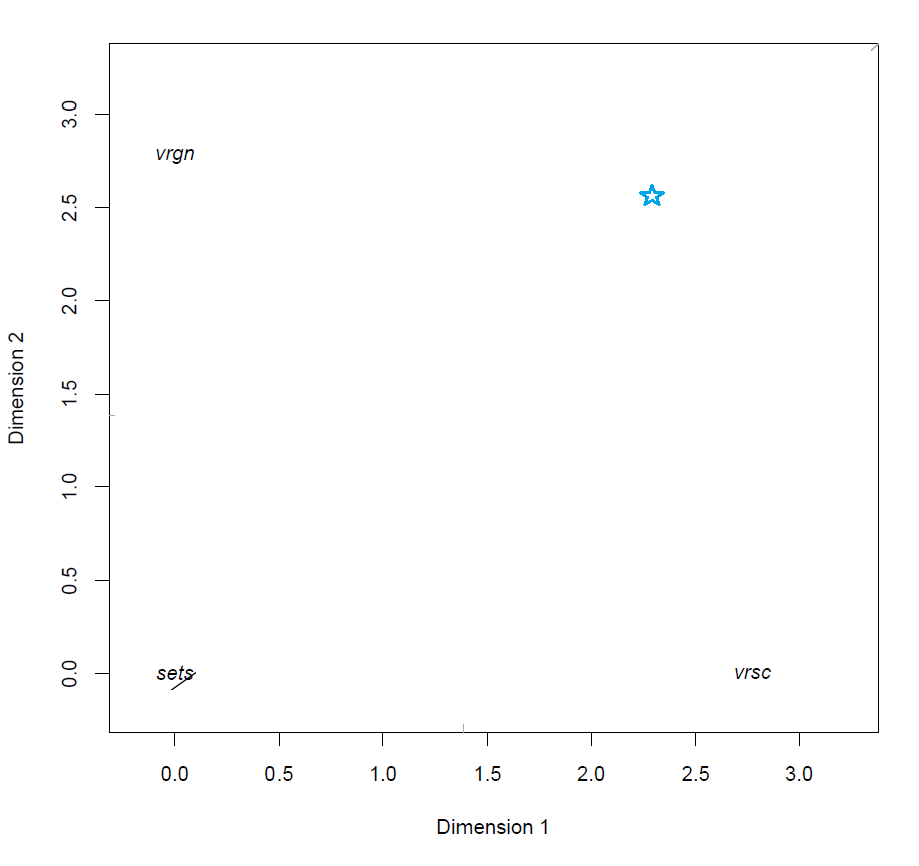
Below is a visual representation of the model fitted on a set of measurements of iris flowers.
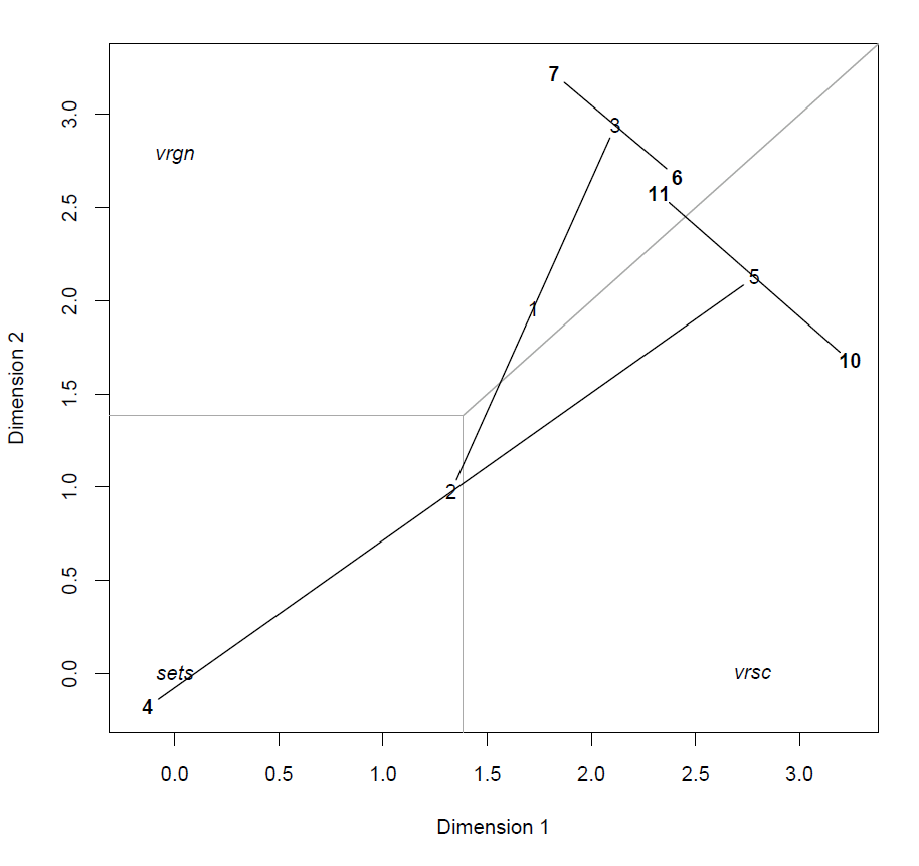
MLRP model of Iris data
We will explain the technique by looking at the various components of the figure above and how these interact in the model.
Model components
The maximum dimensionality is dependent on the number of outcome classes (), such that
. This is in line with statistical technique that rely on a base category approach in which the probability of the classes is estimated with respect to the a specific class( for example logistic regression), making one set of parameters redundant to estimate. In our example of the Iris data we fit a model in maximum dimensionality and therefore the class point do not need to be estimated. It would be possible to reduce the number of dimensions, but we will not go into this now. In our example data we have three different species we want to predict meaning that 2 is the maximum number of dimensions.
Model space
There are 2 key component in the model which determine the class probability of an observation, namely the class point ()and the subject points (
). The former are represented in the figure by their corresponding class label. Because we are creating a model in maximum dimensionality the coordinates of the class points are fixed, for example Setosa (sets) is located at [0, 0].
The subject points are the locations of the subjects (different Iris plants) within the model space. The coordinate for subject on dimension
is computes by:
,
where ranges over all predictors of observation
. In other words, observations
are transformed into
by using the model parameters
. The latter is optimized during the model construction phase. (We‘ ll go into the modeling procedure later on.)
Subject point
The class probability is computed by the relative distance of the subject point and all the class points:
.
denotes a distance function, which we defined as the squared Euclidean
distance between the subject () and class point (
), such
that:
.
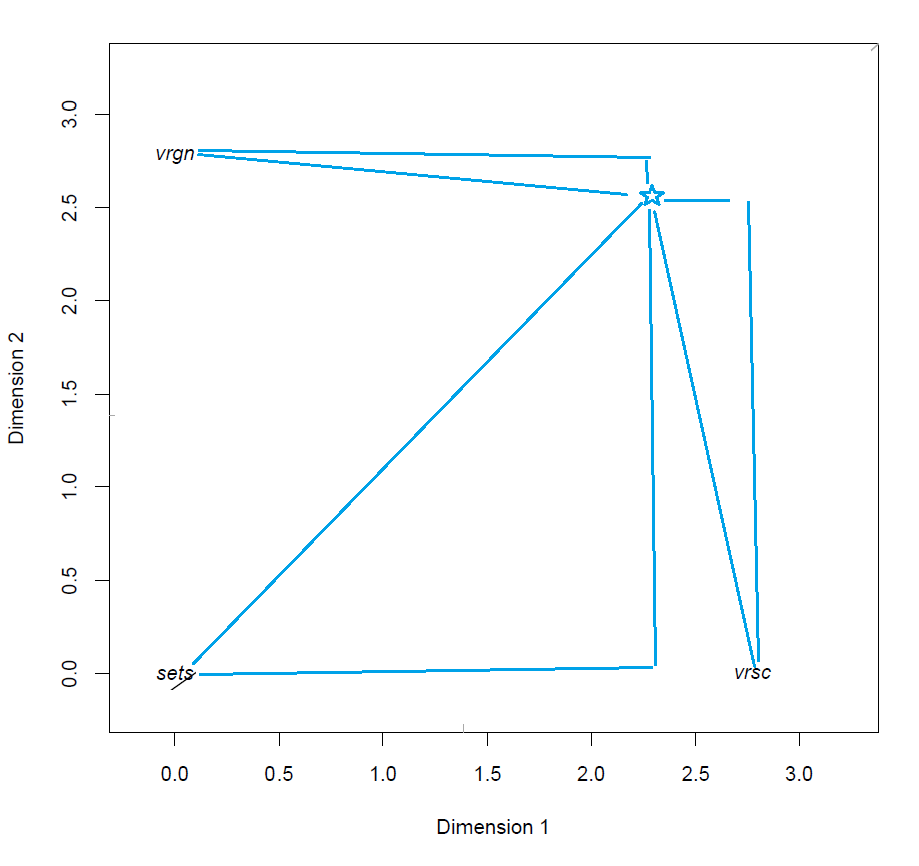
Distance between subject and class points
This means that the predicted class of a subject is the class corresponding to the closest class point of a subject. This is made clear in the model visualization by drawing the gray lines to represent the decision boundaries between the classes, such that subjects which fall within that region have the highest (predicted) probability of belonging to that class.
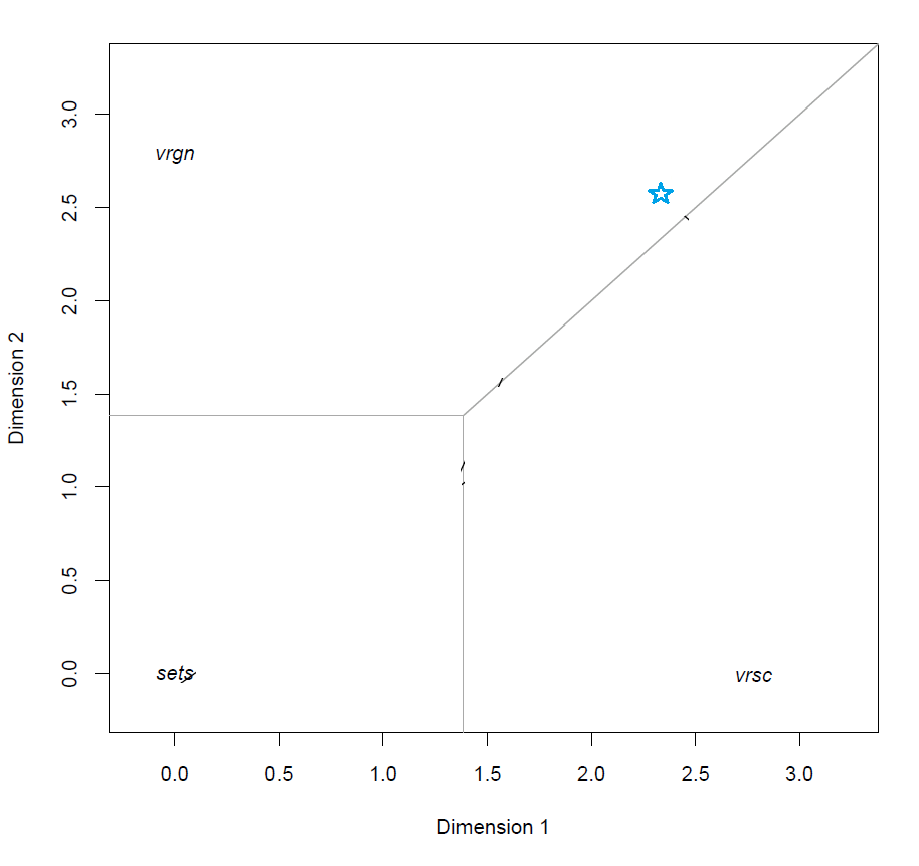
Class regions
Modeling procedure
As described before, the location of the subject is modelled. Originally the subject observations were used, but in this new technique we created sub-groups of observations and project these in the model space. This procedure of partitioning data in order to classify subjects is also used in the construction of decision trees, in which the data is recursively partitioned in sub-groups in order to obtain more homogeneous sets of subjects with respect to the outcome classes.
In this approach, the tree is built such that the split is selected which maximizes the log-likelihood. After a new split is found, and thus new branches are formed, the location of the new nodes is optimized by (re)estimating the regression weights . This procedure is repeated until the number of subjects in the end nodes is to small. In order to select the optimal tree size Akaike information criterion (AIC) is used. A benefit of the AIC is that it is not only using the log-likelihood, but also takes the number of model parameters into account.
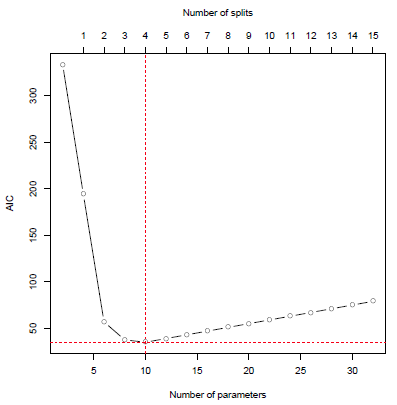
AIC values during model development
In the graph above it can be seen that the AIC value quickly improves in the first few steps. After the optimal value at 4 splits, the performance measure steadily deteriorates as more splits are added to the model. The resulting tree model is represented below.
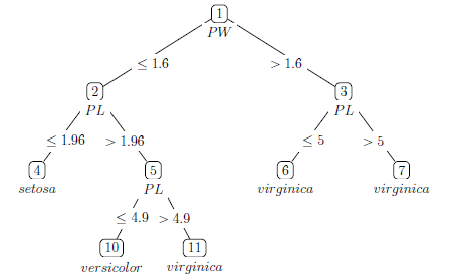
Iris tree model
In order to classify a subject we start at the top: if the petal width 1.6 move to the left branch, if not move to the right branch. Next, let’s go down the left branch here the split is made based on whether the petal length
1.96. This process is repeated until there are no splits anymore, in this case we have ended up in an end/terminal node. The end nodes indicate the predicted outcome class; for example flowers ending up in node 4 are classified as being a Setosa species.
The class membership is estimated based on the distance framework discussed above and the tree can therefore be represented in the model space with the following figure.

MLRP model of Iris data
In this visualization we see that the trunk of the tree is located in the Virginica region. When the first split is evaluated and the pedal width 1.6, the subjects ‘move’ to the Setosa region. Interesting to not is that the second node is very close to the decision boundary with Versicolour, meaning that the probability for this class is also relatively high. The next split (petal length
1.96), provides a big discrimination between these classes, because node 4 is located close to the class point Setosa (and further away from the other classes), whereas node 5 moves away from that class point. We can see that the sub-group ending up in node 5 are split up and either end up in the Versicolour or Virginica region. It might seem that the split at node 3 is redundant, because both child nodes are located in the Virginica region. However the predicted probability is different for each node; node 6 is closer to the decision boundary with Versicolour, meaning that the probability for both classes is more similar.